In-Silico Search Analysis of Potential Inhibitors for 3-Chymotrypsin-Like Protease Of Sars-Cov-2 (Covid-19).
DOI:
https://doi.org/10.37934/jrnn.4.1.4956Keywords:
COVID19, molecular docking, 3CLPROAbstract
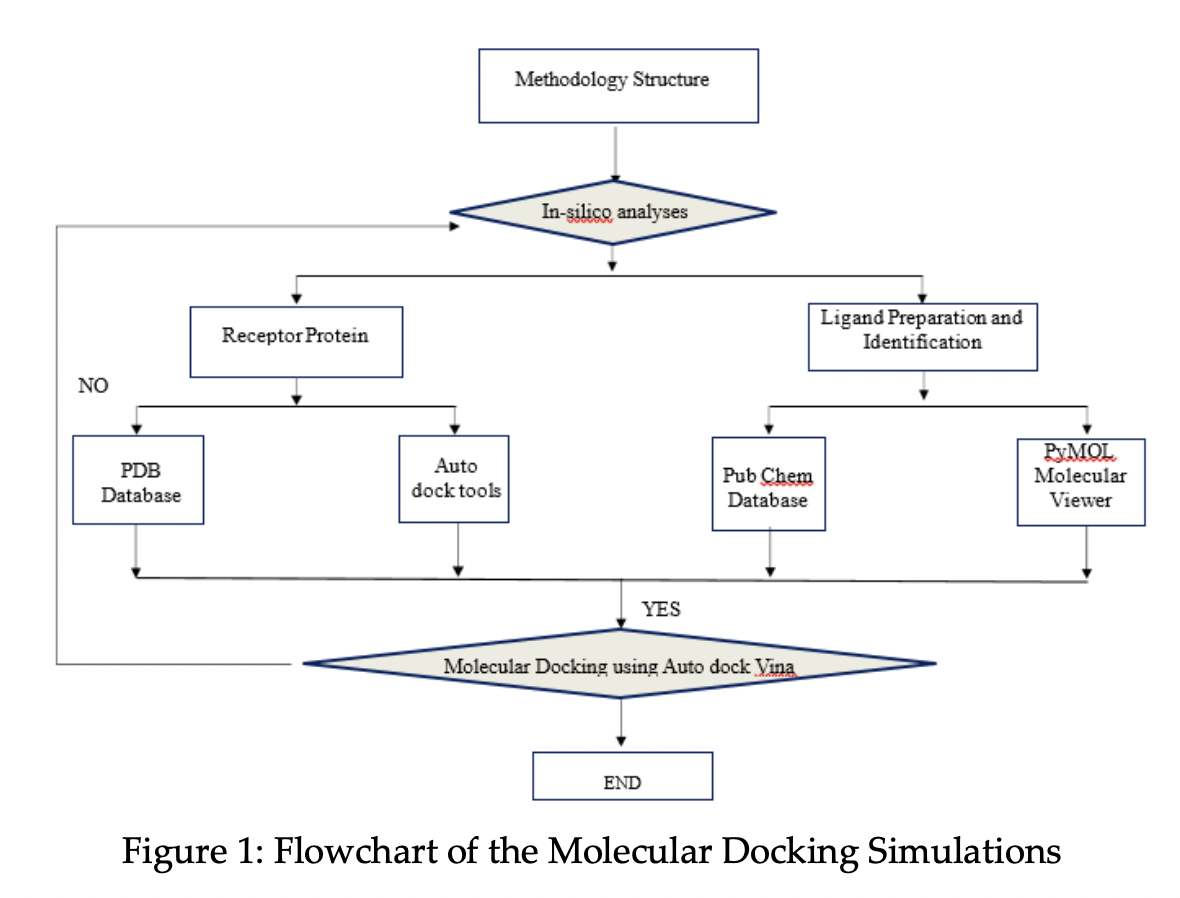
In this study, three commercial drugs, Imatinib, Nilotinib and Dexamethasone were docked against 3 Chymotrypsin-like Protease or the 3CLPRO of the Severe Acute Respiratory Syndrome (SARS-CoV-2) of coronavirus COVID-19 using the program Auto dock Vina. The protein and the ligands were downloaded from the Protein Data Bank and Pub Chem, respectively. The docked conformations were analysed in terms of the molecular interactions such as hydrogen bonds and hydrophobic interactions. The best drug with the lowest binding energy was Nilotinib with -9.5 kcal/mol followed by Imatinib -9.1 kcal/mol. The objective of this study is to investigate the effectiveness of the three commercial drugs against the protein 3CLPRO.
Downloads